Omicron or B.1.1.529, the last “version” of SARS-CoV-2 original strain, was identified almost two months ago.
Today, Omicron is going to replace Delta variant prevalence in almost all countries.
According to ECDC monitor of Gisaid data, at the moment we write, Omicron variant is already prevalent in all European countries, with a distribution percentage close to 60% while Delta, the once-dominant, nowadays is close to 30%.
Different data are coming, instead, from the US: while the variant trend is similar to European countries, Omicron proportion is now near to 100% in almost all States.
Omicron BA.2 variant highly underestimate
Based on its genomic data, Omicron variant has been straight separated into three subgroups: BA.1, BA.2, and BA.3.
These clades share most of their specific mutations except for the deletion used as the target for S Gene Target Failure (SGTF/ S gene dropout) that is not present in BA.2.
The too-quick decision made to adopt the SGTF molecular method to diagnose Omicron has therefore caused the underestimation of the variant cases.
What’s new?
BA.2 cases are spiraling, gradually reaching the numbers of BA.1.
But this is not happening because the variant has reached a greater thrust, but probably because of the underestimation given by the SGTF inadequacy.
SGTF is not valid for defining Omicron circulation: sequencing data have clearly defined this point.
As we have been saying since Omicron first appearance, the only way to make a diagnosis is through a positive detection and not through a negative analysis.
Omicron variants (BA.1, BA.2, and BA.3) must be diagnosed by detecting their own characteristic single-base mutations and not by the absence of a region and therefore the absence of a signal.
Most of the media reporting new BA.2 spreading claim that PCR test failed to detect the variant, but things are different: SGTF test has failed.
Since it’s impossible to sequence all the samples due to a matter of time and costs required, it’s necessary that all the laboratories work to provide diagnosis using PCR tests able to detect specific SARS CoV-2 genes mutations besides not mutated genes (like our Covid VoC Rapid PCR Typing kit). This is the only way not to lose cases, variants, and mutations.
Although there is no evidence that the behavior of the BA.2 clade is different from the others, it is necessary to monitor its progression in a continuous and capillary way in order to not be surprised by the events.
Written by Simone Di Giacomo, PhD, OaCP R&D Manager
Here we go!
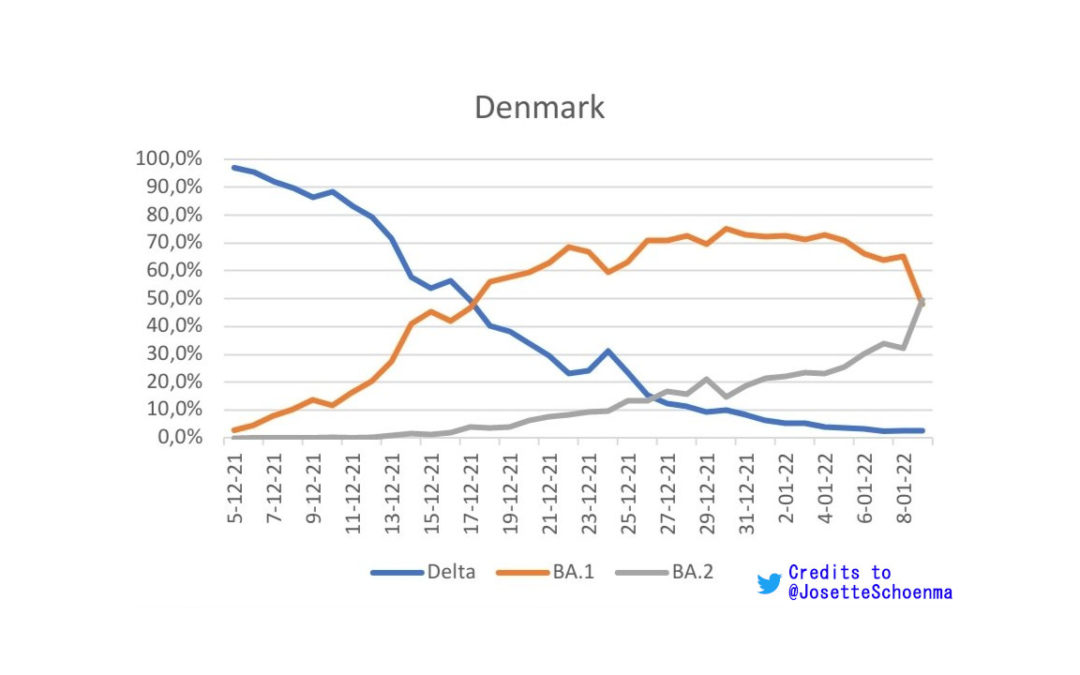
A dominant BA.2 in Denmark on the 9th of January 2022.
2 days later than I predicted and maybe some extra when data will be complete. But the 2k samples available on the 7th, make it quite clear were it was going already, if you ask me.https://t.co/peK0Qpxl9Z pic.twitter.com/wlSXOfBsDN
— Josette Schoenmakers (@JosetteSchoenma) January 14, 2022